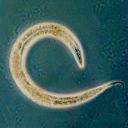
Caenorhabditis elegans is a free-living, transparent nematode, about 1 mm in length, that lives in temperate soil environments. In 1974, Sydney Brenner began research into the molecular and developmental biology of C. elegans, which has since been extensively used as a model organism, being the first multicellular species to have its whole genome sequenced.
Assembly
Genome sequence and annotation have been imported from the WS269 release of WormBase (which includes the WBcel235 version of the C.elegans reference genome). Included are annotated operons, genes, transcripts and translations, as well as RNAi and BLAST/BLAT homology data. In addition Affymetrix and Agilent cross references for C.elegans expression arrays have been added.
We include C.elegans in Ensembl (and also Ensembl Genomes) to allow people to access the data through the Ensembl user interface (both for visualisation and data mining) and to provide cross-species integration through our comparative genomics resources (such as homologous gene links and protein family pages).
Other assemblies
- WS190 (Ensembl release 54)
Gene annotation
This species and other invertebrates are also available from our sister site, Ensembl Metazoa
More information
General information about this species can be found in Wikipedia.
Statistics
Summary
| Assembly | WBcel235, INSDC Assembly GCA_000002985.3, Dec 2012 |
| Base Pairs | 100,286,401 |
| Golden Path Length | 100,286,401 |
| Assembly provider | WormBase |
| Annotation provider | WormBase |
| Annotation method | Import |
| Genebuild started | Jan 2022 |
| Genebuild released | Oct 2014 |
| Genebuild last updated/patched | Oct 2014 |
| Database version | 115.282 |
Gene counts
| Coding genes | 19,985 |
| Non coding genes | 24,813 |
| Small non coding genes | 24,519 |
| Long non coding genes | 294 |
| Pseudogenes | 2,128 |
| Gene transcripts | 60,000 |