Variation Table (Gene)
Short sequence variations are shown by consequence type in the Variation Table.
This view shows all the variant consequences in a gene. If the same variant falls in several transcripts within the same gene, a new row will be displayed for each transcript.
The table columns are as follows:
- ID - The identifier of this particular genetic variant in an external database. Frequently, this will be an rs identifier, a reference SNP from NCBI dbSNP.
- Chr:bp - Chromosome name and base pair co-ordinates
- Alleles - Possible nucleotides at the position listed in the Chr:bp column. These are reported for the forward strand of the genome sequence
- Global MAF - The global minor allele frequency is calculated using all 1000 Genomes Phase 3 data for this SNP, across populations
- Class - For example, Single Nucleotide Polymorphism (SNP) and Insertion-Deletion (InDel)
- Source - The database/project from which the variant was imported
- Evidence Data that support a variant and suggest how reliable the variant is. A summary of evidence status is available on our documentation on Variation data.
- Clin. sig. – A term assigned by dbSNP and ClinVar that indicates pathogenicity or drug response. A list of clinical significance terms can be found on our documentation on Variation data.
- Conseq. type - Consequence type based on SO (Sequence Ontology) terms.
- AA - (Amino acid) Possible amino acid(s) (if any) resulting from the allele(s) at the position of the variation. Data is only shown for missense variants.
- AA coord - (Amino acid coordinates) The position of the amino acid (AA) within the protein sequence.
- SIFT - Prediction of variant effect on protein function by SIFT
- PolyPhen - Prediction of variant effect on protein function by PolyPhen
- CADD - Prediction of variant effect on protein function by CADD
- REVEL - Prediction of variant effect on protein function by REVEL
- MetaLR - Prediction of variant effect on protein function by MetaLR
- MutationAssessor - Prediction of variant effect on protein function by MutationAssessor
- Transcript - The specific Ensembl transcript for the gene of interest in which the variant shows the consequence type
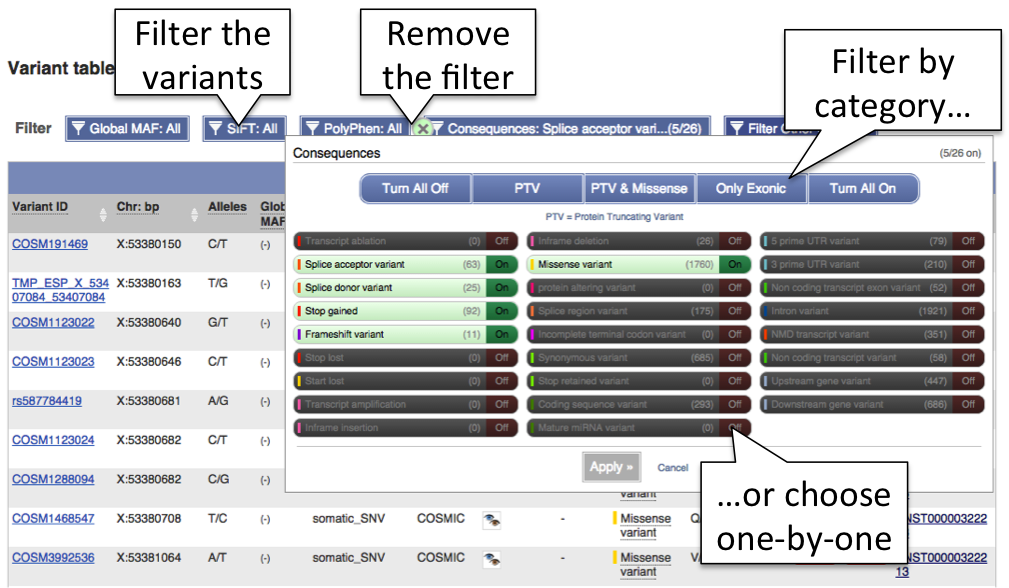
You can order the table by the columns by clicking on the up/down arrows by the column titles. Filters above the table allow you to filter the data by various parameters.